Publications
Papers are listed below. * denote joint first authors.
2026
-
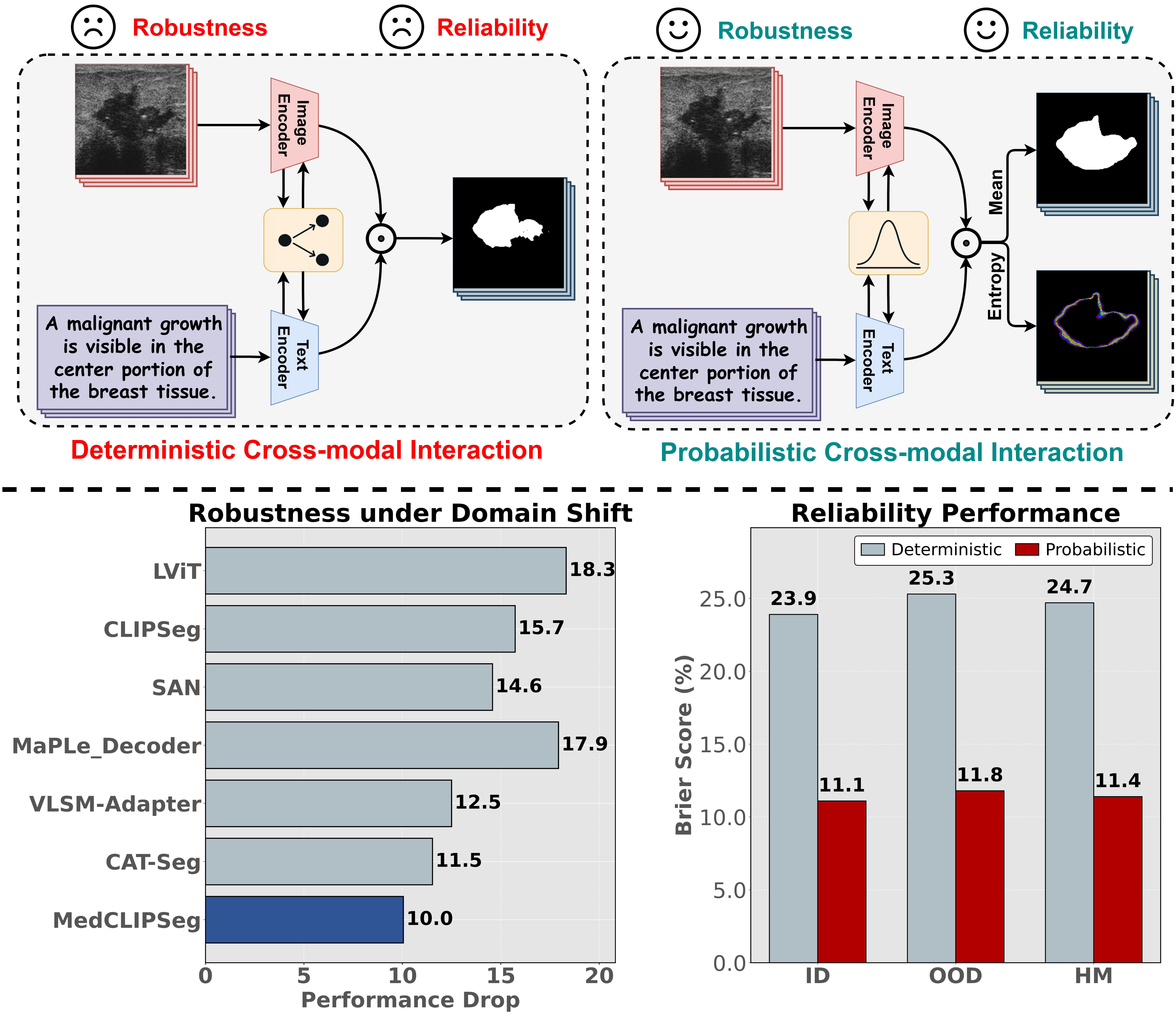 MedCLIPSeg: Probabilistic Vision–Language Adaptation for Data-Efficient and Generalizable Medical Image SegmentationTaha Koleilat, Hojat Asgariandehkordi, Omid Nejati Manzari, Berardino Barile, Yiming Xiao*, and Hassan Rivaz*In Proceedings of the Computer Vision and Pattern Recognition Conference, 2026
MedCLIPSeg: Probabilistic Vision–Language Adaptation for Data-Efficient and Generalizable Medical Image SegmentationTaha Koleilat, Hojat Asgariandehkordi, Omid Nejati Manzari, Berardino Barile, Yiming Xiao*, and Hassan Rivaz*In Proceedings of the Computer Vision and Pattern Recognition Conference, 2026Medical image segmentation remains challenging due to limited annotations for training, ambiguous anatomical features, and domain shifts. While vision-language models such as CLIP offer strong cross-modal representations, their potential for dense, text-guided medical image segmentation remains underexplored. We present MedCLIPSeg, a novel framework that adapts CLIP for robust, data-efficient, and uncertainty-aware medical image segmentation. Our approach leverages patch-level CLIP embeddings through probabilistic cross-modal attention, enabling bidirectional interaction between image and text tokens and explicit modeling of predictive uncertainty. Together with a soft patch-level contrastive loss that encourages more nuanced semantic learning across diverse textual prompts, MedCLIPSeg effectively improves data efficiency and domain generalizability. Extensive experiments across 16 datasets spanning five imaging modalities and six organs demonstrate that MedCLIPSeg outperforms prior methods in accuracy, efficiency, and robustness, while providing interpretable uncertainty maps that highlight local reliability of segmentation results. This work demonstrates the potential of probabilistic vision-language modeling for text-driven medical image segmentation.
@inproceedings{koleilat2026medclipseg, title = {MedCLIPSeg: Probabilistic Vision–Language Adaptation for Data-Efficient and Generalizable Medical Image Segmentation}, author = {Koleilat, Taha and Asgariandehkordi, Hojat and Nejati Manzari, Omid and Barile, Berardino and Xiao*, Yiming and Rivaz*, Hassan}, booktitle = {Proceedings of the Computer Vision and Pattern Recognition Conference}, year = {2026}, } -
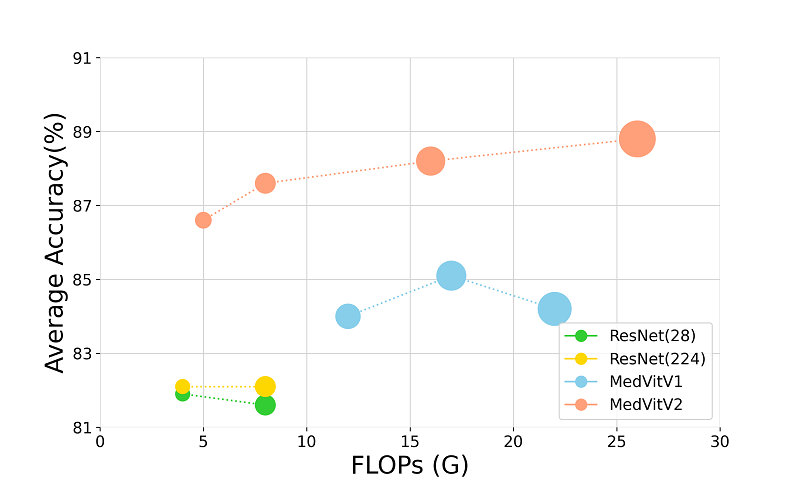 Medical image classification with KAN-integrated transformers and dilated neighborhood attentionOmid Nejati Manzari, Hojat Asgariandehkordi, Taha Koleilat, Yiming Xiao, and Hassan RivazApplied Soft Computing, 2026
Medical image classification with KAN-integrated transformers and dilated neighborhood attentionOmid Nejati Manzari, Hojat Asgariandehkordi, Taha Koleilat, Yiming Xiao, and Hassan RivazApplied Soft Computing, 2026@article{NEJATIMANZARI2026114045, title = {Medical image classification with KAN-integrated transformers and dilated neighborhood attention}, journal = {Applied Soft Computing}, volume = {186}, pages = {114045}, year = {2026}, issn = {1568-4946}, doi = {https://doi.org/10.1016/j.asoc.2025.114045}, url = {https://www.sciencedirect.com/science/article/pii/S1568494625013584}, author = {{Nejati Manzari}, Omid and Asgariandehkordi, Hojat and Koleilat, Taha and Xiao, Yiming and Rivaz, Hassan}, }
2025
-
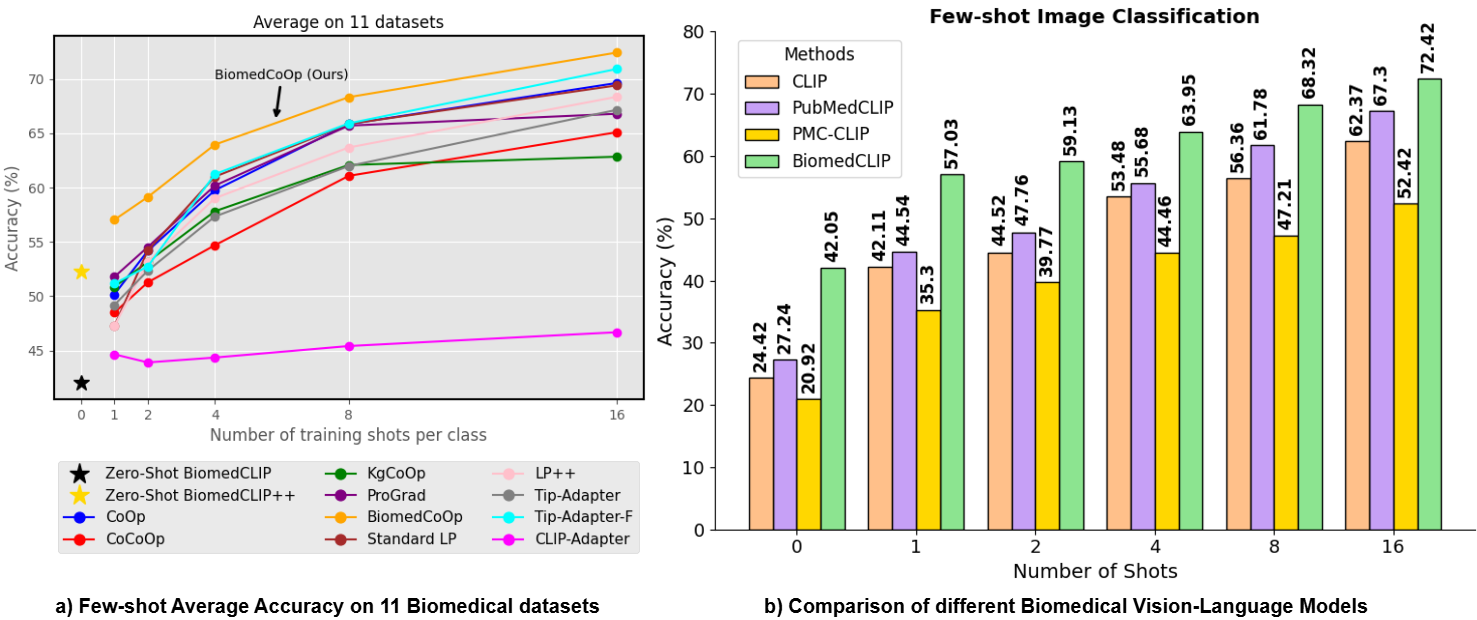 Biomedcoop: Learning to prompt for biomedical vision-language modelsTaha Koleilat, Hojat Asgariandehkordi, Hassan Rivaz, and Yiming XiaoIn Proceedings of the Computer Vision and Pattern Recognition Conference, 2025
Biomedcoop: Learning to prompt for biomedical vision-language modelsTaha Koleilat, Hojat Asgariandehkordi, Hassan Rivaz, and Yiming XiaoIn Proceedings of the Computer Vision and Pattern Recognition Conference, 2025Recent advancements in vision-language models (VLMs), such as CLIP, have demonstrated substantial success in self-supervised representation learning for vision tasks. However, effectively adapting VLMs to downstream applications remains challenging, as their accuracy often depends on time-intensive and expertise-demanding prompt engineering, while full model fine-tuning is costly. This is particularly true for biomedical images, which, unlike natural images, typically suffer from limited annotated datasets, unintuitive image contrasts, and nuanced visual features. Recent prompt learning techniques, such as Context Optimization (CoOp) intend to tackle these issues, but still fall short in generalizability. Meanwhile, explorations in prompt learning for biomedical image analysis are still highly limited. In this work, we propose BiomedCoOp, a novel prompt learning framework that enables efficient adaptation of BiomedCLIP for accurate and highly generalizable few-shot biomedical image classification. Our approach achieves effective prompt context learning by leveraging semantic consistency with average prompt ensembles from Large Language Models (LLMs) and knowledge distillation with a statistics-based prompt selection strategy. We conducted comprehensive validation of our proposed framework on 11 medical datasets across 9 modalities and 10 organs against existing state-of-the-art methods, demonstrating significant improvements in both accuracy and generalizability.
@inproceedings{koleilat2025biomedcoop, title = {Biomedcoop: Learning to prompt for biomedical vision-language models}, author = {Koleilat, Taha and Asgariandehkordi, Hojat and Rivaz, Hassan and Xiao, Yiming}, booktitle = {Proceedings of the Computer Vision and Pattern Recognition Conference}, pages = {14766--14776}, year = {2025}, } -
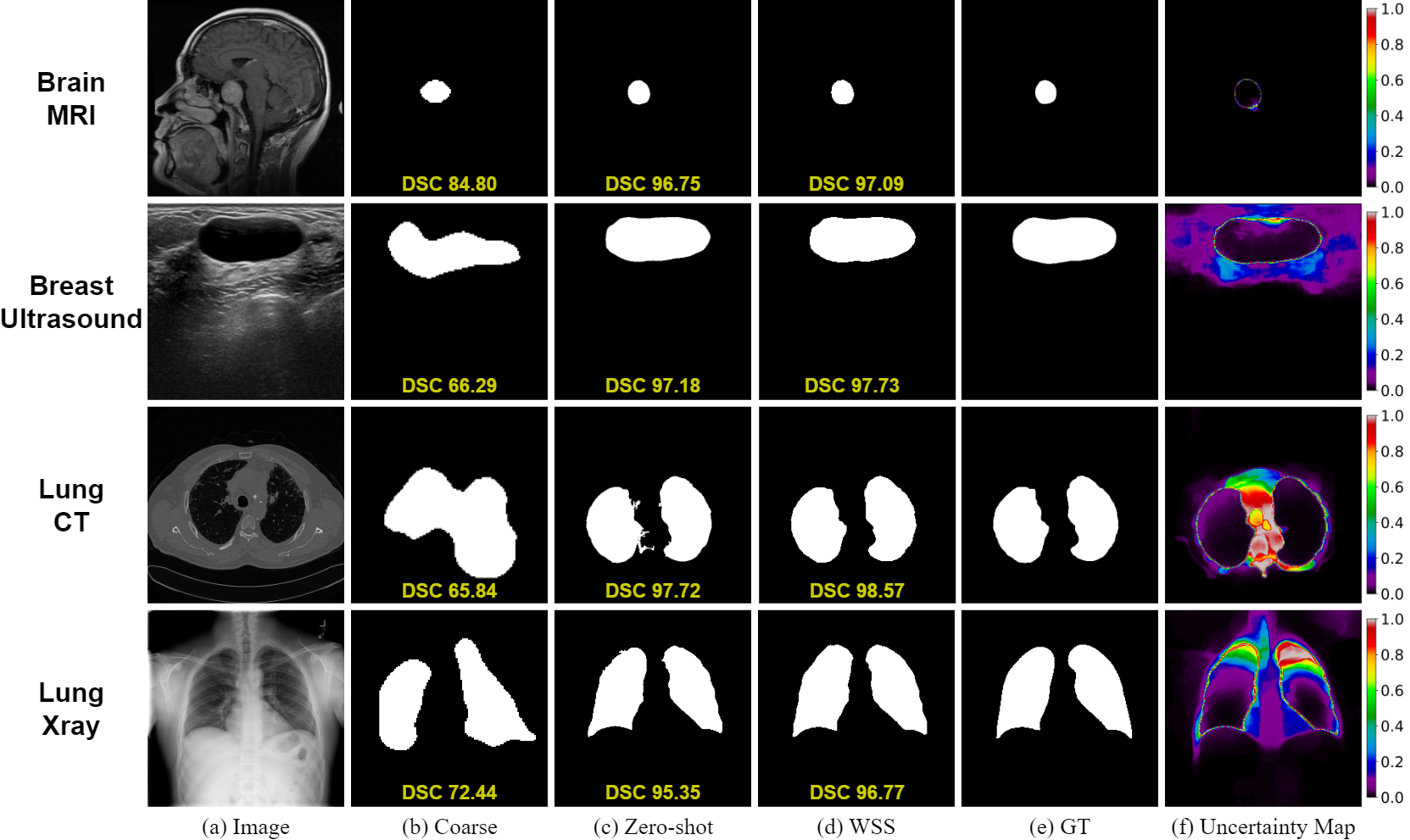 Medclip-samv2: Towards universal text-driven medical image segmentationTaha Koleilat, Hojat Asgariandehkordi, Hassan Rivaz, and Yiming XiaoMedical Image Analysis, 2025
Medclip-samv2: Towards universal text-driven medical image segmentationTaha Koleilat, Hojat Asgariandehkordi, Hassan Rivaz, and Yiming XiaoMedical Image Analysis, 2025Segmentation of anatomical structures and pathologies in medical images is essential for modern disease diagnosis, clinical research, and treatment planning. While significant advancements have been made in deep learning-based segmentation techniques, many of these methods still suffer from limitations in data efficiency, generalizability, and interactivity. Recently, foundation models like CLIP and Segment-Anything-Model (SAM) have paved the way for interactive and universal image segmentation. In this work, we introduce MedCLIP-SAMv2, a framework that integrates BiomedCLIP and SAM to perform text-driven medical image segmentation in zero-shot and weakly supervised settings. The approach fine-tunes BiomedCLIP with a new DHN-NCE loss and leverages M2IB to create visual prompts for SAM; we also explore uncertainty-aware refinement via checkpoint ensembling.
@article{koleilat2025medclip, title = {Medclip-samv2: Towards universal text-driven medical image segmentation}, author = {Koleilat, Taha and Asgariandehkordi, Hojat and Rivaz, Hassan and Xiao, Yiming}, journal = {Medical Image Analysis}, pages = {103749}, year = {2025}, publisher = {Elsevier}, } -
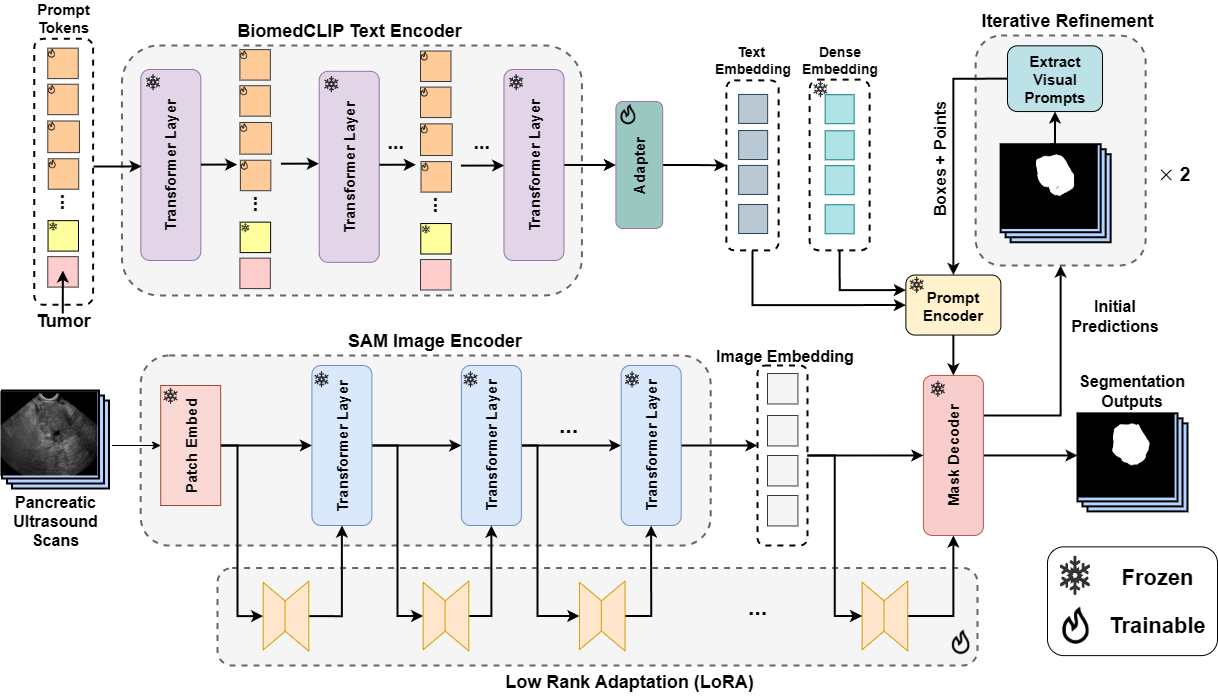 Textsam-eus: Text prompt learning for sam to accurately segment pancreatic tumor in endoscopic ultrasoundPascal Spiegler*, Taha Koleilat*, Arash Harirpoush, Corey S Miller, Hassan Rivaz, Marta Kersten-Oertel, and Yiming XiaoIn Proceedings of the IEEE/CVF International Conference on Computer Vision, 2025
Textsam-eus: Text prompt learning for sam to accurately segment pancreatic tumor in endoscopic ultrasoundPascal Spiegler*, Taha Koleilat*, Arash Harirpoush, Corey S Miller, Hassan Rivaz, Marta Kersten-Oertel, and Yiming XiaoIn Proceedings of the IEEE/CVF International Conference on Computer Vision, 2025Pancreatic cancer carries a poor prognosis and relies on endoscopic ultrasound (EUS) for targeted biopsy and radiotherapy. However, the speckle noise, low contrast, and unintuitive appearance of EUS make segmentation of pancreatic tumors with fully supervised deep learning (DL) models both error-prone and dependent on large, expert-curated annotation datasets. To address these challenges, we present TextSAM-EUS, a novel, lightweight, text-driven adaptation of the Segment Anything Model (SAM) that requires no manual geometric prompts at inference. Our approach leverages text prompt learning (context optimization) through the BiomedCLIP text encoder in conjunction with a LoRA-based adaptation of SAM’s architecture to enable automatic pancreatic tumor segmentation in EUS, tuning only 0.86% of the total parameters. On the public Endoscopic Ultrasound Database of the Pancreas, TextSAM-EUS with automatic prompts attains 82.69% Dice and 85.28% normalized surface distance (NSD), and with manual geometric prompts reaches 83.10% Dice and 85.70% NSD, outperforming both state-of-the-art (SOTA) supervised DL models and foundation models (e.g., SAM and its variants). As the first attempt to incorporate prompt learning in SAM-based medical image segmentation, TextSAM-EUS offers a practical option for efficient and robust automatic EUS segmentation.
@inproceedings{spiegler2025textsam, title = {Textsam-eus: Text prompt learning for sam to accurately segment pancreatic tumor in endoscopic ultrasound}, author = {Spiegler*, Pascal and Koleilat*, Taha and Harirpoush, Arash and Miller, Corey S and Rivaz, Hassan and Kersten-Oertel, Marta and Xiao, Yiming}, booktitle = {Proceedings of the IEEE/CVF International Conference on Computer Vision}, pages = {948--957}, year = {2025}, } -
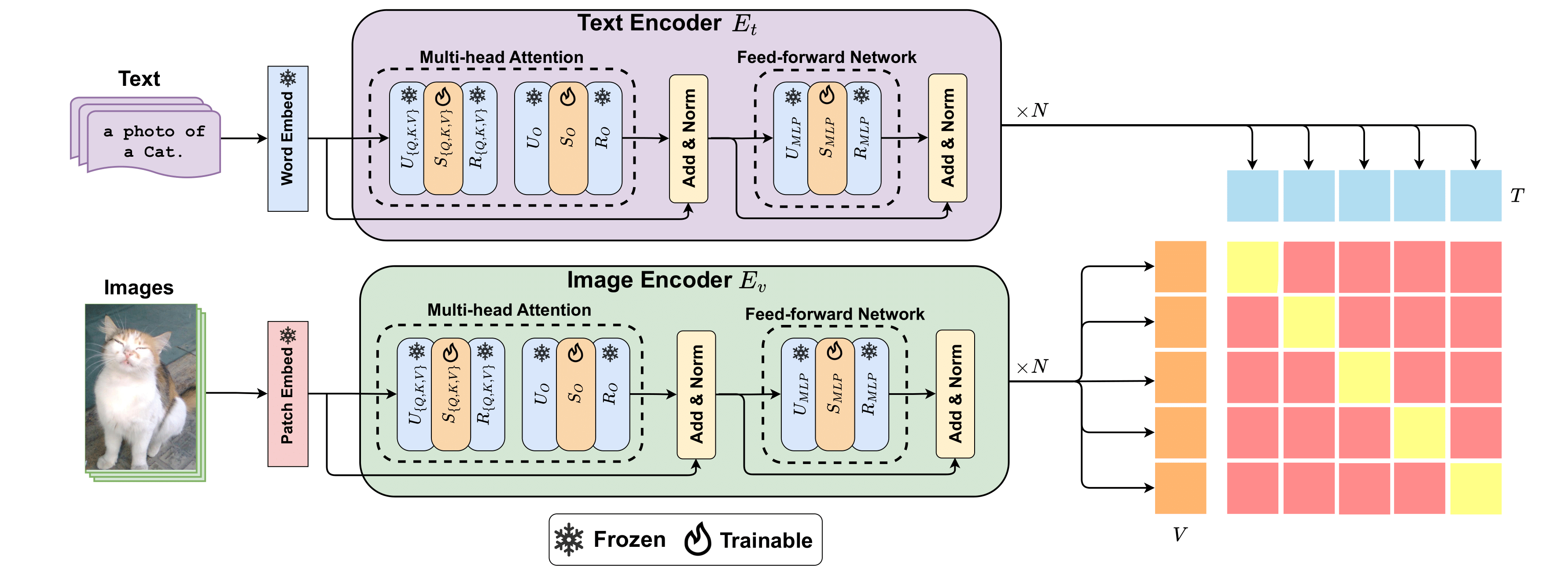 Singular Value Few-shot Adaptation of Vision-Language ModelsTaha Koleilat, Hassan Rivaz, and Yiming XiaoarXiv preprint arXiv:2509.03740, 2025
Singular Value Few-shot Adaptation of Vision-Language ModelsTaha Koleilat, Hassan Rivaz, and Yiming XiaoarXiv preprint arXiv:2509.03740, 2025Vision-language models (VLMs) like CLIP have shown impressive zero-shot and few-shot learning capabilities across diverse applications. However, adapting these models to new fine-grained domains remains difficult due to reliance on prompt engineering and the high cost of full model fine-tuning. Existing adaptation approaches rely on augmented components, such as prompt tokens and adapter modules, which could limit adaptation quality, destabilize the model, and compromise the rich knowledge learned during pretraining. In this work, we present CLIP-SVD, a novel multi-modal and parameter-efficient adaptation technique that leverages Singular Value Decomposition (SVD) to modify the internal parameter space of CLIP without injecting additional modules. Specifically, we fine-tune only the singular values of the CLIP parameter matrices to rescale the basis vectors for domain adaptation while retaining the pretrained model. This design enables enhanced adaptation performance using only 0.04% of the model’s total parameters and better preservation of its generalization ability. CLIP-SVD achieves state-of-the-art classification results on 11 natural and 10 biomedical datasets, outperforming previous methods in both accuracy and generalization under few-shot settings. Additionally, we leverage a natural language-based approach to analyze the effectiveness and dynamics of the CLIP adaptation to enable interpretability of CLIP-SVD.
@article{koleilat2025singular, title = {Singular Value Few-shot Adaptation of Vision-Language Models}, author = {Koleilat, Taha and Rivaz, Hassan and Xiao, Yiming}, journal = {arXiv preprint arXiv:2509.03740}, year = {2025}, } -
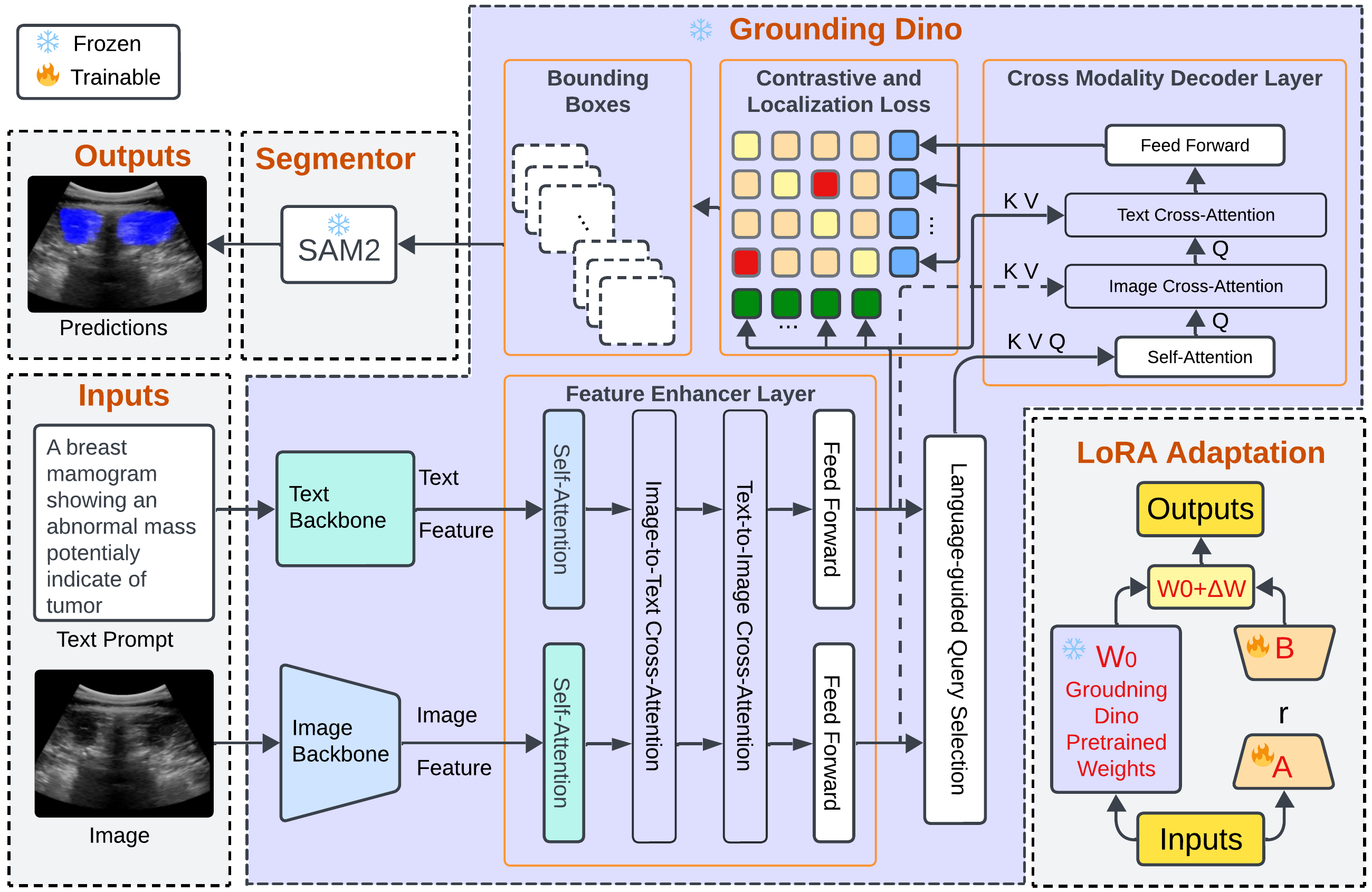 Grounding DINO-US-SAM: Text-Prompted Multiorgan Segmentation in Ultrasound With LoRA-Tuned Vision–Language ModelsHamza Rasaee, Taha Koleilat, and Hassan RivazIEEE Transactions on Ultrasonics, Ferroelectrics, and Frequency Control, 2025
Grounding DINO-US-SAM: Text-Prompted Multiorgan Segmentation in Ultrasound With LoRA-Tuned Vision–Language ModelsHamza Rasaee, Taha Koleilat, and Hassan RivazIEEE Transactions on Ultrasonics, Ferroelectrics, and Frequency Control, 2025@article{11146904, author = {Rasaee, Hamza and Koleilat, Taha and Rivaz, Hassan}, journal = {IEEE Transactions on Ultrasonics, Ferroelectrics, and Frequency Control}, title = {Grounding DINO-US-SAM: Text-Prompted Multiorgan Segmentation in Ultrasound With LoRA-Tuned Vision–Language Models}, year = {2025}, volume = {72}, number = {10}, pages = {1414-1425}, keywords = {Ultrasonic imaging;Image segmentation;Breast;Grounding;Training;Imaging;Adaptation models;Acoustics;Thyroid;Liver;Grounding DINO;prompt-driven segmentation;segment anything model (SAM) SAM2;ultrasound image segmentation;vision–language models (VLMs)}, doi = {10.1109/TUFFC.2025.3605285}, }
2024
-
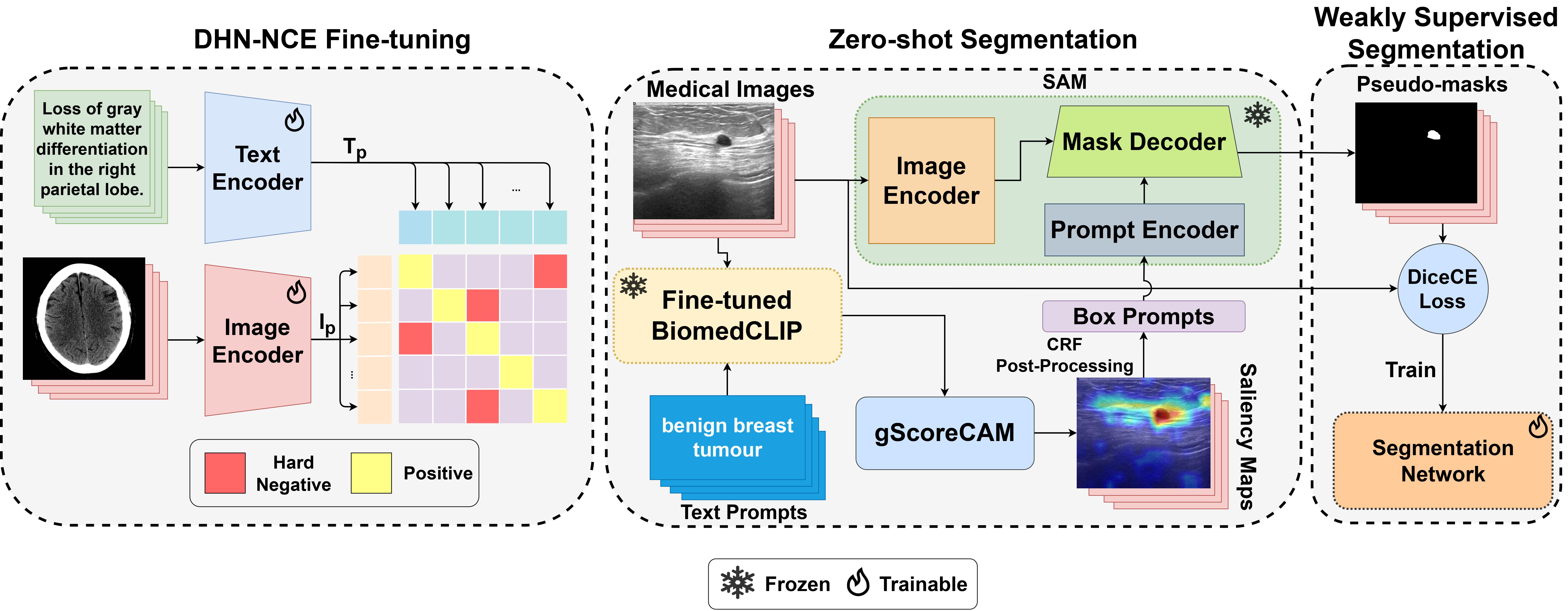 Medclip-sam: Bridging text and image towards universal medical image segmentationTaha Koleilat, Hojat Asgariandehkordi, Hassan Rivaz, and Yiming XiaoIn International conference on medical image computing and computer-assisted intervention, 2024
Medclip-sam: Bridging text and image towards universal medical image segmentationTaha Koleilat, Hojat Asgariandehkordi, Hassan Rivaz, and Yiming XiaoIn International conference on medical image computing and computer-assisted intervention, 2024Medical image segmentation of anatomical structures and pathology is crucial in modern clinical diagnosis, disease study, and treatment planning. While deep learning-based methods have achieved strong performance, they often lack data efficiency, generalizability, and interactability. In this work, we propose MedCLIP-SAM, a novel framework that bridges vision–language models and segmentation foundation models to enable text-driven universal medical image segmentation. MedCLIP-SAM integrates BiomedCLIP fine-tuned with a Decoupled Hard Negative Noise Contrastive Estimation (DHN-NCE) loss, gScoreCAM-based saliency generation, CRF post-processing, and Segment Anything Model (SAM) refinement. The framework supports both zero-shot and weakly supervised segmentation and is validated across breast ultrasound, brain MRI, and chest X-ray datasets, demonstrating strong accuracy and generalization.
@inproceedings{koleilat2024medclip, title = {Medclip-sam: Bridging text and image towards universal medical image segmentation}, author = {Koleilat, Taha and Asgariandehkordi, Hojat and Rivaz, Hassan and Xiao, Yiming}, booktitle = {International conference on medical image computing and computer-assisted intervention}, pages = {643--653}, year = {2024}, organization = {Springer}, }